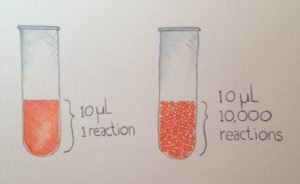
 Two new papers from the McCarroll lab describe ways of using nanoliter droplets to answer questions in genetics and biology. Droplets provide a way of scaling molecular biological reactions across tens of thousands of tiny reaction compartments.
Two new papers from the McCarroll lab describe ways of using nanoliter droplets to answer questions in genetics and biology. Droplets provide a way of scaling molecular biological reactions across tens of thousands of tiny reaction compartments.
In Drop-phase, Jack Regan, Nolan Kamitaki and colleagues describe a way to quickly determine the chromosomal phase of multiple sequence variants, even when those variants are separated by substantial genomic distances. Their approach involves partitioning genomic DNA across thousands of droplets, then analyzing how alleles from different loci co-partition across the droplets.
In Drop-seq, Evan Macosko and colleagues describe a way to profile genome-wide gene expression in thousands of individual cells, in facile, inexpensive experiments. The approach involves enacpsulating cells in droplets while using beads to deliver distinct DNA barcodes to each droplet. Macosko and colleagues used the approach to profile 44,808 individual cells from the mouse retina.